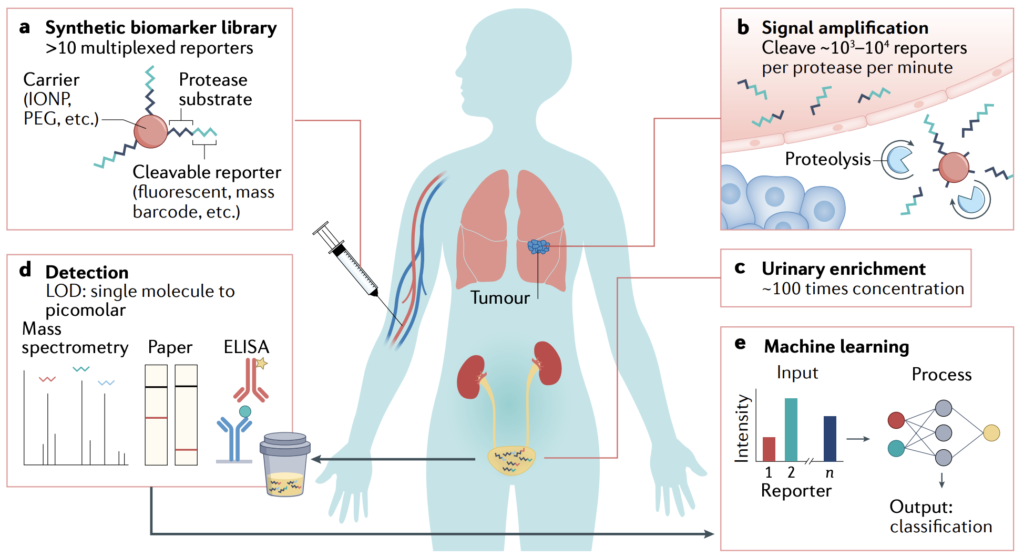
BME researchers combine precision and simplicity in transforming diagnostic tools.
Georgia Tech researchers have developed biosensors with advanced sleuthing skills and the technology may revolutionize cancer detection and monitoring.
The tiny detectives can identify key biological markers using logical reasoning inspired by the “AND” function in computers — like, when you need your username and password to log in. And unlike traditional biosensors comprised of genetic materials — cells, bits of DNA — these are made of manufactured molecules.
These new biosensors are more precise and simpler to manufacture, reducing the number of false positives and making them more practical for clinical use. And because the sensors are cell-free, there’s a reduced risk for immunogenic side effects.
“We think the accuracy and simplicity of our biosensors will lead to accessible, personalized, and effective treatments, ultimately saving lives,” said Gabe Kwong, associate professor and Robert A. Milton Endowed Chair in the Wallace H. Coulter Department of Biomedical Engineering, who led the study, published this month in Nature Nanotechnology.
Breaking With Tradition
The researchers set out to address the limitations in current biosensors for cancer, like the ones designed for CAR-T cells to allow them to recognize tumor cells. These advanced biosensors are made of genetic material, and there is growing interest to reduce the potential for off-target toxicity by using Boolean “AND-gate” computer logic. That means they’re designed to release a signal only when two specific conditions are met.
“Traditionally, these biosensors involve genetic engineering using cell-based systems, which is a complex, time-consuming, and expensive process,” said Kwong.
So, his team developed biosensors made of iron oxide nanoparticles and special molecules called cyclic peptides. Synthesizing nanomaterials and peptides is a simpler, less costly process than genetic engineering, according to Kwong, “which means we can likely achieve large-scale, economical production of high-precision biosensors.”
Unlocking the AND-gate
Biosensors detect cancer signals and track treatment progress by turning biological signals into readable outputs for doctors. With AND-gate logic, two distinct inputs are required for an output.
Accordingly, the researchers engineered cyclic peptides — small amino acid chains — to respond only when they encounter two specific types of enzymes, proteases called granzyme B (secreted by the immune system) and matrix metalloproteinase (from cancer cells). The peptides generate a signal when both proteases are present and active.
Think of a high-security lock that needs two unique keys to open. In this scenario, the peptides are the lock, activating the sensor signal only when cancer is present and being confronted by the immune system.
“Our peptides allow for greater accuracy in detecting cancer activity,” said the study’s lead author, Anirudh Sivakumar, a postdoctoral researcher in Kwong’s Laboratory for Synthetic Immunity. “It’s very specific, which is important for knowing when immune cells are targeting and killing tumor cells.”
Super Specific
In animal studies, the biosensors successfully distinguished between tumors that responded to a common cancer treatment called immune checkpoint blockade therapy — ICBT, which enhances the immune system — from tumors that resisted treatment.
During these tests, the sensors also demonstrated their ability to avoid false signals from other, unrelated health issues, such as when the immune system confronted a flu infection in the lungs, away from the tumor.
“This level of specificity can be game changing,” Kwong said. “Imagine being able to identify which patients are responding to the therapy early in their treatment. That would save time and improve patient outcomes.”
The first step toward this simpler, precise form of cancer diagnostics began with an ambitious but humble ($50,000) seed grant from the Petit Institute for Bioengineering and Bioscience five years ago for a collaboration between Kwong’s lab and the lab of M.G. Finn, professor and chair in the School of Chemistry and Biochemistry.
It evolved into a multi-institutional project supported by grants from the National Science Foundation and National Institutes of Health that included researchers from the University of California-Riverside, as well as Georgia Tech faculty researchers Finn and Peng Qiu, associate professor in the Coulter Department.
“The progression of the research, from an initial seed grant all the way to animal studies, was very smooth,” Kwong said. “Ultimately, a collaborative, multidisciplinary effort turned our early vision into something that could have a great impact in healthcare.”
Citation: Anirudh Sivakumar, Hathaichanok Phuengkham, Hitha Rajesh, Quoc D. Mac, Leonard C. Rogers, Aaron D. Silva Trenkle, Swapnil Subhash Bawage, Robert Hincapie, Zhonghan Li, Sofia Vainikos, Inho Lee, Min Xue, Peng Qiu, M. G. Finn, Gabriel A. Kwong. “AND-gated protease-activated nanosensors for programmable detection of anti-tumour immunity.” Nature Nanotechnology (January 2025). https://doi.org/10.1038/s41565-024-01834-8
Funding: This research was supported in part by National Institutes of Health (NIH) grants 5U01CA265711, 5R01CA237210, 1DP2HD091793, and 5DP1CA280832.
Contact
Jerry Grillo
Communications
Wallace H. Coulter Department of Biomedical Engineering